library(planex)
#> Loading required package: dplyr
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
#> Loading required package: ggplot2
library(tidyverse)
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ lubridate 1.9.5 ✔ tibble 3.3.1
#> ✔ purrr 1.2.2 ✔ tidyr 1.3.2
#> ✔ readr 2.2.0
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
library(planex)
library(tidyverse)
data(rendimento2k)
glimpse(rendimento2k)
#> Rows: 12
#> Columns: 3
#> $ rendimento <int> 28, 36, 18, 31, 25, 32, 19, 30, 27, 32, 23, 29
#> $ reagente <factor2k> -1, 1, -1, 1, -1, 1, -1, 1, -1, 1, -1, 1
#> $ catalisador <factor2k> -1, -1, 1, 1, -1, -1, 1, 1, -1, -1, 1, 1
fit <- aov(rendimento ~ reagente*catalisador, data=rendimento2k)
testResiduals(fit)
#>
#> Shapiro-Wilk normality test
#>
#> data: resid
#> W = 0.906, p-value = 0.1895
#>
#> ------------------------------------------
#> Bartlett test of Homogeneity of Variances:
#> Bartlett's K-squared df p.value
#> reagente 0.169806150 1 0.6802841
#> catalisador 0.002058164 1 0.9638148
#>
#> -----------------------------------------------
#> Durbin-Watson Test for Autocorrelated Errors:
#> lag Autocorrelation D-W Statistic p-value
#> 1 -0.2978723 2.507092 0.276
#> Alternative hypothesis: rho != 0
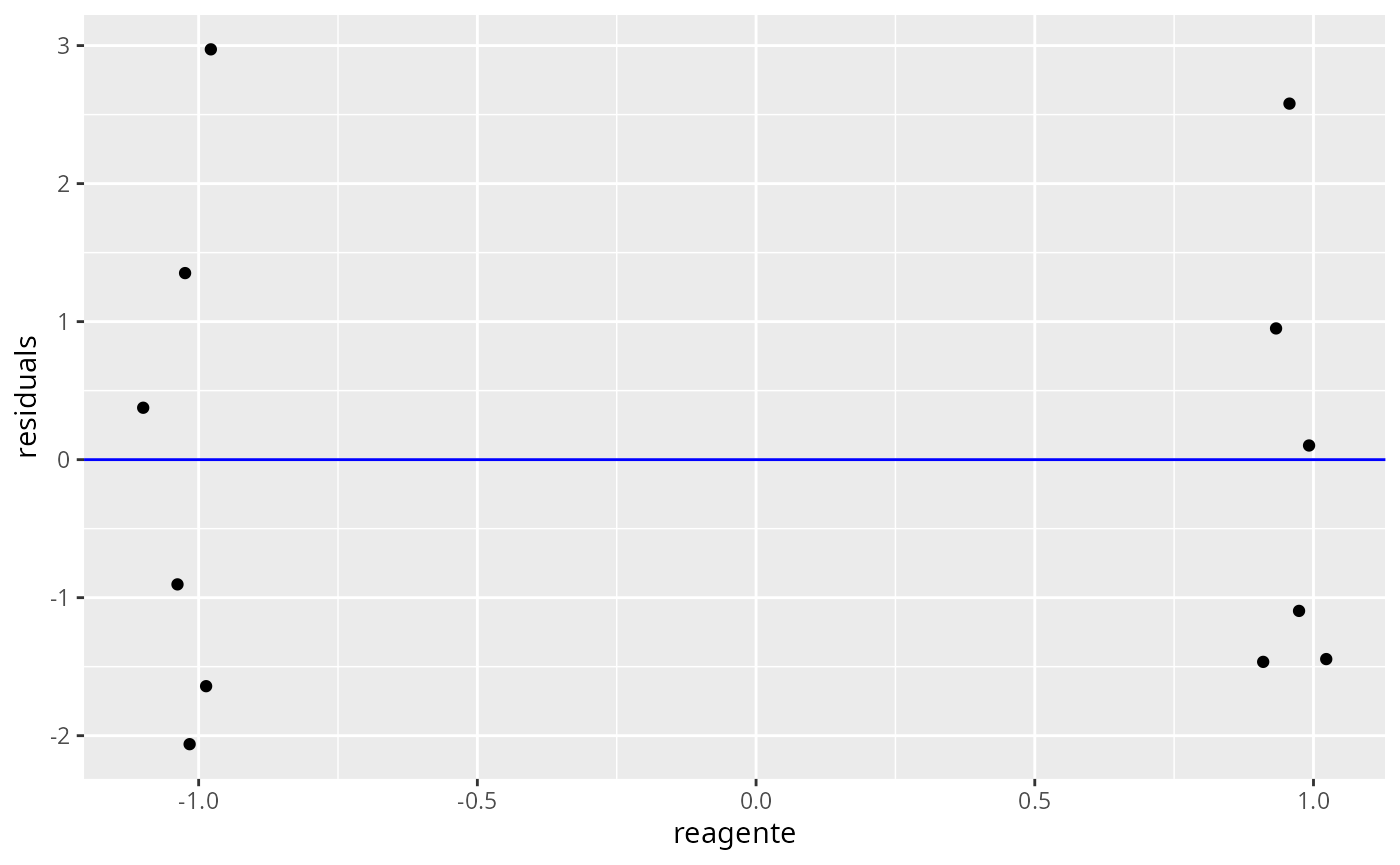
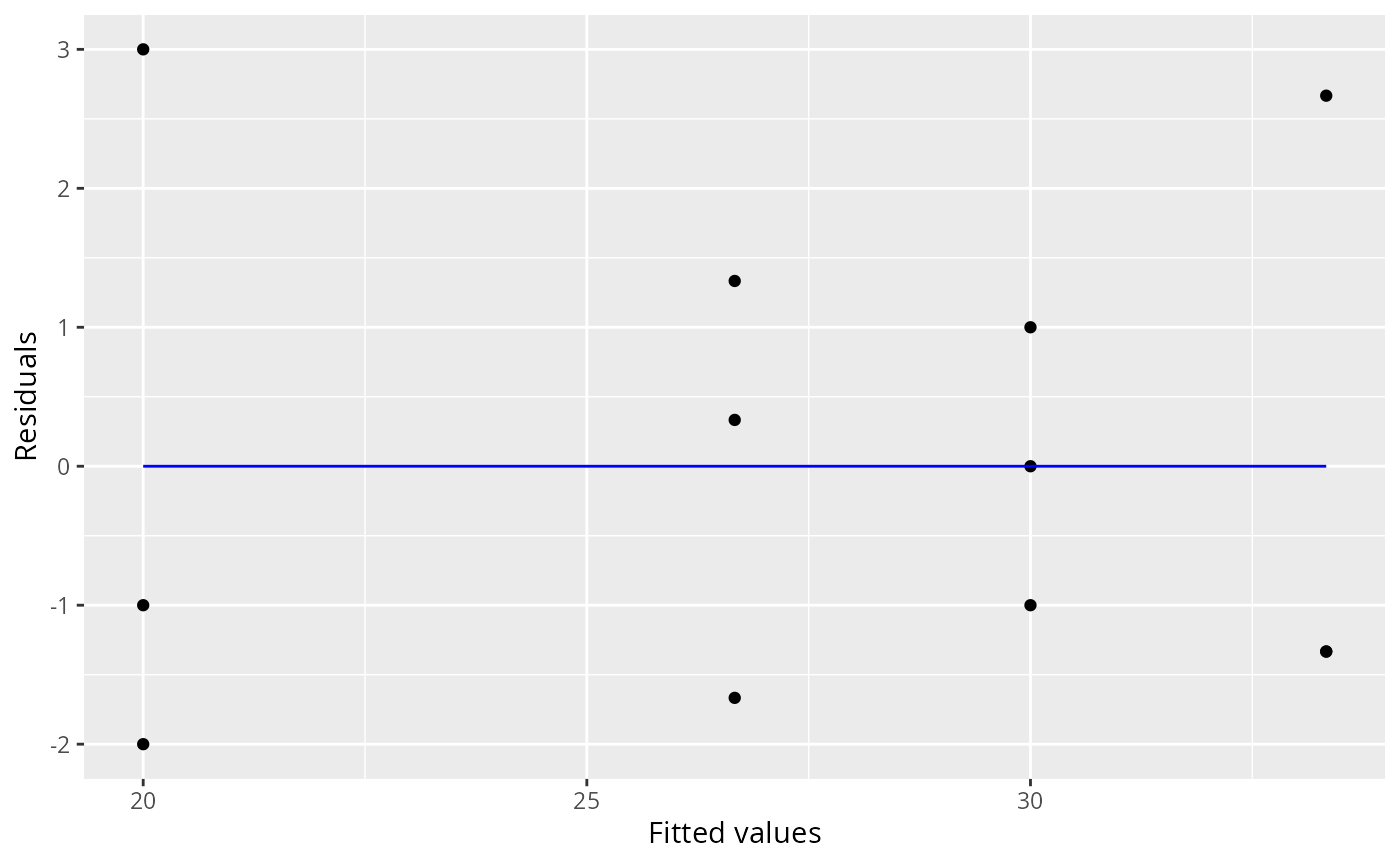
ggresiduals(fit)
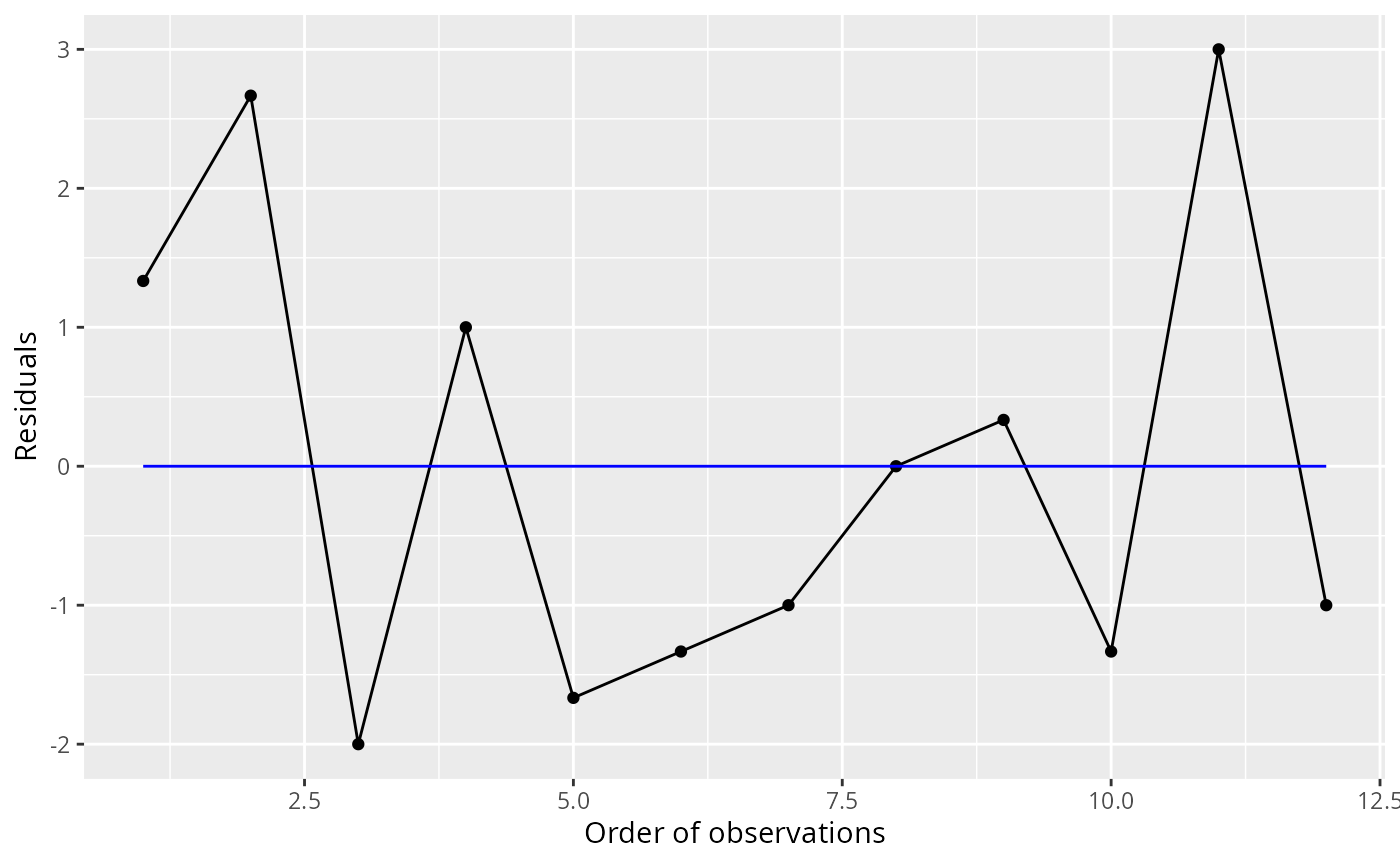
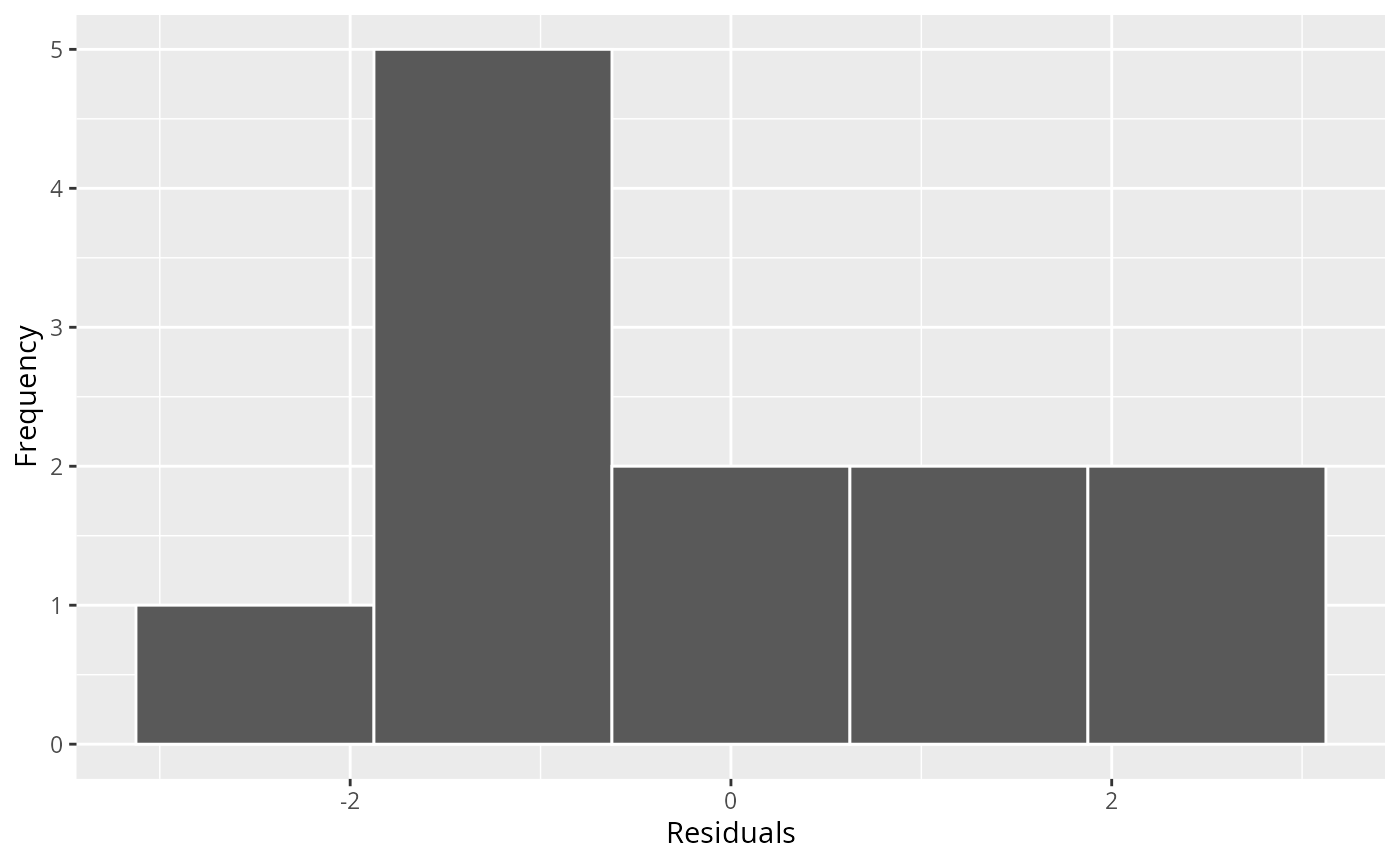
#> `geom_smooth()` using method = 'loess' and formula = 'y ~ x'
#> Warning in simpleLoess(y, x, w, span, degree = degree, parametric = parametric,
#> : pseudoinverse used at 19.933
#> Warning in simpleLoess(y, x, w, span, degree = degree, parametric = parametric,
#> : neighborhood radius 10.067
#> Warning in simpleLoess(y, x, w, span, degree = degree, parametric = parametric,
#> : reciprocal condition number 5.7985e-17
#> Warning in simpleLoess(y, x, w, span, degree = degree, parametric = parametric,
#> : There are other near singularities as well. 45.338
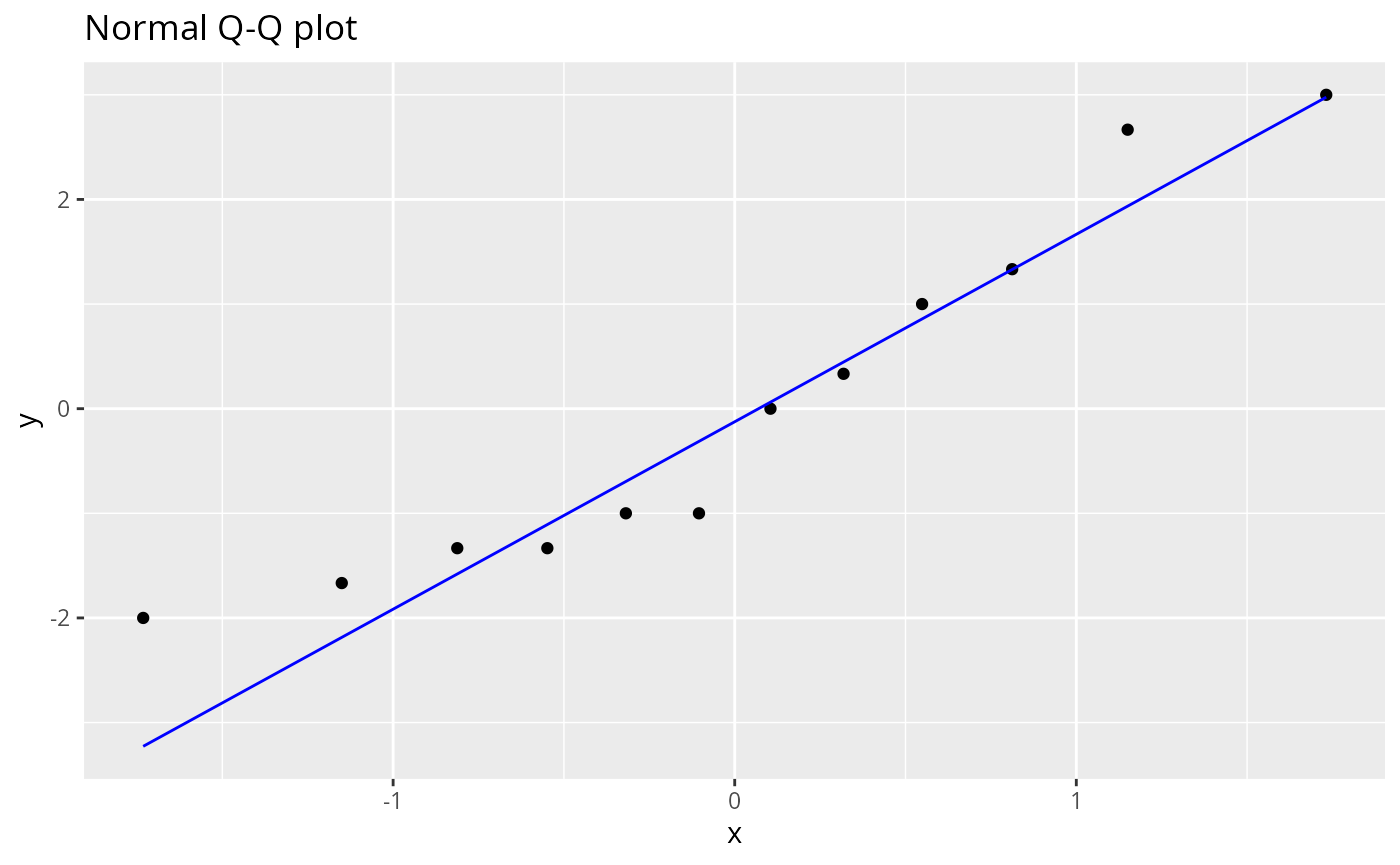
#> Don't know how to automatically pick scale for object of type <factor2k>.
#> Defaulting to continuous.
#> Don't know how to automatically pick scale for object of type <factor2k>.
#> Defaulting to continuous.